The database represents genomes assembled to different levels: Viral genomes are included if they are in the NCBI Reference Sequence (RefSeq) database or have been selected as viral neighbor genomes by the NCBI viral genomes group ( more details). Similarly, plasmids are only included when they are associated with chromosome sequences. Organelle genomes are included only when there is also a nuclear genome assembly. The Assembly resource includes prokaryotic and eukaryotic genomes with a Whole Genome Shotgun (WGS) assembly, clone-based assembly, or completely sequenced genome (gapless chromosomes). DDBJ, ENA or GenBank, and the assembly represented in the NCBI Reference Sequence (RefSeq) project. It also tracks the relationship between an assembly submitted to the International Nucleotide Sequence Database Collaboration ( INSDC ), i.e. The web resource provides meta-data about assemblies such as assembly names (and alternate names), simple statistical reports of the assembly (type and number of contigs, scaffolds N50s) and a history view of updates.
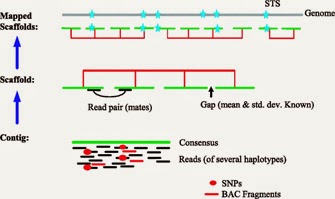
The database provides a versioned Assembly accession number that tracks changes to assemblies as they are updated by submitting groups over time. The Assembly database has information about the structure of assembled genomes as represented in an AGP file or as a collection of completely sequenced chromosomes. Information presented for each assembly.

